Professor Senjie Lin is profiled on the Member Spotlight of the American Society for the Advancement of Science (AAAS). His research on dinoflagellate biology and its relevance to addressing harmful algal blooms, coral bleaching, and other climate change challenges is highlighted. The Spotlight, which can be found in https://www.aaas.org/membership/member-spotlight/senjie-lin-explores-potential-hidden-answers-tiny-dinoflagellates, also reveals how Lin was drawn to science in general and to dinoflagellate work in particular.
Author: Schuler, Debra
When Life Gives You Lemons … Hold a Virtual International Fish Conference
By Elaina Hancock.
UConn Marine Sciences researcher Hannes Baumann left the 2019 Larval Fish Conference, the 43rd installment of the annual conference, with excitement for the 2020 American Fisheries Society Larval Fish Conference which was to be hosted by UConn. The planning and overall experience has been entirely different than he expected and turned out to be a real “lemons to lemonade” situation, says Baumann.
“For over 40 years, this small but important gathering has happily meandered between North American and European locations. The last in-person conference was 2019 in Palma de Mallorca. A treat. For 2020, I agreed that it now was my turn to organize a Larval Fish Conference. We were so excited, but you know what happened next,” says Baumann.
The 2020 conference had to be canceled outright, and plans started for 2021 in hopes it could be held in-person, but those too were later changed.
“We canceled the June 2020 in-person conference, but naively only postponed it by one year to June 2021, thinking then that one year later, we surely would be done with this virus and all the travel restrictions. So much for that. Again, in March of this year, we had to cancel the 2021 in-person meeting, but replaced it with a three-day virtual meeting that myself and a team organized in the months leading up to the conference.”
The ability to shift gears and ensure forward momentum is a valuable skill the workforce quickly acquired as a result of the pandemic. Baumann says the amount of help and hard work provided by University Events and Conference Services staff to make the switch to virtual has been essential.
“The staff has done a fantastic job with this. You can’t imagine how many things can go wrong, but the staff have solved all of these problems. We all have such gratitude for all their help.”
In pulling everything together, Baumann says the process was both exhausting and rewarding as turnout exceeded expectations.
“The virtual nature of the meeting led to a record diversity of registrants. The conferences were always international, but that meant largely European and North American countries, plus Japan, Australia and New Zealand. This year, we had talks and posters from 28 different countries, a record.”
Technology made it possible: the WebEx platform in particular, coupled with another platform called Gatherly to encourage networking, a difficult-to-replicate experience for virtual meetings.
“Here I was sitting in my New England office on Gatherly and a video chat would pop up with a colleague in British Columbia, when all of a sudden a little avatar joined in and popped up a conversation from New Zealand. Another person from Europe joined after that, and we were talking like we are talking right now. That was so cool, and the participants loved that,” says Baumann.
Happy with how everything turned out, Baumann says, “The top reaction is the technology gods were smiling on us as there were no major glitches and we are very, very, happy for that. I’m a little surprised that everything went so well.”
Benefits of the virtual conference included increased diversity of participants, many who may not have been able to participate otherwise, a reduced carbon footprint, and participants being able to see more talks, since all were recorded. However, Baumann says meeting in-person still can’t be beat.
“This pandemic is particularly hard for early career researchers, like grad students who want to share their research. It’s important to start talking about their research and be in front of people talking about their work. Though responses have been positive, all participants agree on the fact that even the best run virtual conference cannot replace the quality of networking or personal contact that an in-person meeting can deliver.”
Going forward, Baumann says efforts will be made to ensure future conferences have some mixture of both approaches to make sure prospective attendees have their needs met.
“Some participants were really frank in saying they would never be able to attend an in-person meeting far from their own countries. Some could not easily afford the fee for the virtual conference, but we were able to help. Anybody who says, ‘Well, now that we have virtual conferences, the world is on an equal playing field,’ is not seeing the reality that much of the world has not the same resources and internet connectivity than we do here in an institution like UConn.”
Baumann says this is something organizers will have to think carefully about to calibrate, but going forward, scientific conferences may never be quite the same.
Marine Animals Could Be Used to Clean Up Nature’s Big Pollutant: Microplastics
Over the next four years, faculty from the School of Engineering and College of Liberal Arts and Sciences, including marine sciences professors Evan Ward and George McManus, will use a $2 million grant from the National Science Foundation’s Emerging Frontiers in Research and Innovation (EFRI) program to study the use of mussels (bivalves), combined with microplastic-degrading bacteria, to remove microplastics from the discharge of wastewater treatment plants. For more information about this project see https://today.uconn.edu/2021/03/how-marine-animals-could-be-used-to-clean-up-natures-big-pollutant-microplastics/.
Red Tide Prey Defense is a Costly Business
Postdoctoral investigator Gihong Park and Professor Hans Dam published a study in the Proceedings of the Royal Society B that demonstrates a fitness cost of defense in a red tide dinoflagellate. Organisms must defend themselves against their consumers. It has long been hypothesized that defenses such as toxin production may come at a cost in the form of reduced growth. Yet, demonstrating such costs of defense is challenging. Park and Dam’s study presents a novel approach using a growth-related gene to show that when a red tide dinoflagellate (phytoplankton) is exposed to a copepod grazer, it increases toxin production but decreases its growth gene marker, indicating a fitness cost of toxin production. While costly, the defense is adaptive because it lowers the consumer ingestion rate and it allows the dinoflagellate to persist. The findings have important implications for understanding the factors that control the rise and fall of red tide blooms. Such blooms plague coastal regions wreaking havoc on local fisheries economies and threatening public health.
Link to the paper: https://royalsocietypublishing.org/doi/10.1098/rspb.2020.2480
Viruses, the “good”, not the bad or ugly
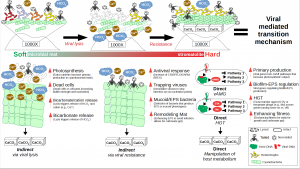
The viruses may be the missing link in the evaluation of life. Formation of Earth’s oldest ecosystems, stromatolites, requires a deeper understanding of several virus-mediated mechanisms that change the cyanobacterial behavior through geologic time, including the “invention” of oxygen-producing photosynthesis. Cyanobacteria also precipitate and cement the carbonate minerals, a stromatolite is preserved in the fossil record, some for as long as 3,500 million years. Without the interception of viruses, this may not have happened and further evolution of life and biogeochemistry that resulted in the Cambrian explosion would have been different. A planet without humans….?
https://today.uconn.edu/2021/02/lifes-surprising-debt-to-viruses/
Mechanisms proposed for virus-cyanobacteria interaction. Image: Richard White III
Marine Sciences graduate student Alec Shub working with Governor’s Council on Climate Change
For the past several months Alec Shub has been working with Connecticut’s Department of Energy and Environmental Protection (DEEP), coordinating reports for the Governor’s Council on Climate Change (GC3). The GC3 was established by executive order, in an effort to mitigate greenhouse gas emissions and address strategies for adaptation and resilience to the impacts of climate change throughout the state of Connecticut. One of the Council’s goals is to help the state meet its target of an 80% reduction in greenhouse gas emissions (below 2001 levels) by 2050. This effort has garnered collaboration between an eclectic team of 23 council members and 230 working group members, including experts from a wide range of scientific disciplines, state agencies, local governments, non-profits, and businesses. As a result of his work, Alec has learned how different fields are able to come together and collaborate on a central issue. As a graduate student, Alec volunteered with the Connecticut Institute for Resilience and Climate Adaptation (CIRCA), which was an important stepping stone to the GC3 position. He also believes the GC3 experience will be invaluable preparation for his upcoming Knauss Fellowship with the National Oceanic and Atmospheric Administration’s (NOAA) Climate Program Office.
If you are interested in learning more about the GC3 or reading any of the working group reports, you can visit the page on the DEEP website:
https://portal.ct.gov/DEEP/Climate-Change/GC3/Governors-Council-on-Climate-Change
Christening the Automated Larval Fish Rearing System (ALFiRiS) at the DMS Rankin Lab
Rankin Lab, December 2020. After using and tinkering with our experimental system at the Rankin Seawater Lab of the University of Connecticut’s Department of Marine Sciences for over 5 years, it’s finally time to give the baby a name – ALFiRiS.
Over the past years, the Evolutionary Fish Ecology Lab of Prof. Baumann has built an Automated Larval Fish Rearing System (ALFiRiS) to conduct factorial experiments on the climate sensitivity of fishes. It consists of a 3 x 3 array of recirculating units (600L/150gal) that have independent computer-control over their temperature, oxygen, and pH conditions. We use a self-developed LabView (National Instruments) platform to sequentially monitor tank conditions via industrial-grade oxygen and pH sensors (Hach) and then control gas solenoids (air, N2, CO2) to maintain and modulate environmental conditions. The system can apply static as well as fluctuating conditions on diel and tidal scales. Computerized temperature control further allows simulating heatwaves and other non-static thermal regimes. We’ve only begun to explore all of ALFiRiS’ capabilities.
To learn more, go to https://befel.marinesciences.uconn.edu/alfiris/
Dierssen Lab Earns Recognition from NASA
Dr. Heidi Dierssen, Professor in Marine Sciences, and her postdoc Brandon Russell were among the individuals recognized by NASA during its 2020 Honor Awards event that was held on December 1. They were part of the Coral Reef Airborne Laboratory Mission Team that collected and delivered unprecedented data about reef environments.
Living in an anoxic world: Arsenic cycling supports life for billion of years
Elaina Hancock – UConn Communications
Much of life on planet Earth today relies on oxygen to exist, but before oxygen was present on our blue planet, lifeforms likely used arsenic instead. These findings are detailed in research published today in Communications Earth and Environment.
A key component of the oxygen cycle is where plants and some types of bacteria essentially take sunlight, water, and CO2, and convert them to carbohydrates and oxygen, which are then cycled and used by other organisms that breathe oxygen. This oxygen serves as a vehicle for electrons, gaining and donating electrons as it powers through the metabolic processes. However, for half of the time life has existed on Earth, there was no oxygen present, and for the first 1.5 billion years, we really don’t how these systems worked, says lead author of the study and UConn Professor of Marine Sciences and Geosciences Pieter Visscher.
Light-driven, photosynthetic organisms appear in the fossil record as layered carbonate rocks called stromatolites dating to around 3.7 billion years ago, says Visscher. Stromatolite mats are deposited over the eons by microbial ecosystems, with each layer holding clues about life at that time. There are contemporary examples of microbes that photosynthesize in the absence of oxygen using a variety of elements to complete the process, however it’s unclear how this happened in the earliest life forms.
Theories as to how life’s processes functioned in the absence of oxygen have mostly relied on hydrogen, sulfur, or iron as the elements that ferried electrons around to fulfill the metabolic needs of organisms.
As Visscher explains, these theories are contested; for example, photosynthesis is possible with iron, but researchers do not find evidence of that in the fossil record before oxygen appeared some 2.4 billion years ago. Hydrogen is mentioned, yet the energetics and competition for hydrogen between different microbes shows it is highly unfeasible.
Arsenic is another theoretical possibility, and evidence for that was found in 2008. Visscher says the link with arsenic was strengthened in 2014 when he and colleagues found evidence of arsenic-based photosynthesis in deep time. To further support their theory, the researchers needed to find a modern analog to study the biogeochemistry and element cycling.
Finding an analog to the conditions on early Earth is a challenge for a number of reasons, besides the fact that oxygen is now abundant. For instance, the evidence shows early microbes captured atmospheric carbon and produced organic matter at a time when volcanic eruptions were frequent, UV light was intense in the absence of the ozone layer, and oceans were essentially a toxic soup.
Another challenging aspect of working within the fossil record, especially those as ancient as some stromatolites, is that there are few left due to the cycling of rock as continents move. However, a breakthrough happened when the team discovered an active microbial mat, currently existing in the harsh conditions in Laguna La Brava in the Atacama Desert in Chile.
The mats have not been studied previously but present an otherworldly set of conditions, like those of early Earth. The mats are in a unique environment which leaves them in a permanent oxygen-free state at high altitude where they are exposed to wild, daily temperature swings, and high UV conditions. The mats serve as powerful and informative tools for truly understanding life in the conditions of early Earth.
Visscher explains, “We started working in Chile, where I found a blood red river. The red sediments are made up by anoxogenic photosynthetic bacteria. The water is very high in arsenic as well. The water that flows over the mats contains hydrogen sulfide that is volcanic in origin and it flows very rapidly over these mats. There is absolutely no oxygen.”
The team also showed that the mats were making carbonate deposits and creating a new generation of stromatolites. The carbonate materials also showed evidence for arsenic cycling – that arsenic is serving as a vehicle for electrons — proving that the microbes are actively metabolizing arsenic, much like oxygen in modern systems. Visscher says these findings, along with the fossil evidence, gives a strong sense of the early conditions of Earth.
“Arsenic-based life has been a question in terms of, does it have biological role or is it just a toxic compound?” says Visscher.
That question appears to be answered: “I have been working with microbial mats for about 35 years or so. This is the only system on Earth where I could find a microbial mat that worked absolutely in the absence of oxygen.”
Visscher points out that an important tool they used to perform this research is similar to one onboard the Mars Perseverance rover, currently en route to Mars.
“In looking for evidence of life on Mars, they will be looking at iron and probably they should be looking at arsenic also.”
This work was supported by grants from NSF grant OCE 1561173, ISITE project UB18016-BGS-IS and the São Paulo Research Foundation FAPESP, grant 2015/16235-2. You can also find out more about this work in The Conversation.
DMS researchers show fish to grow slower under future oceanic CO2 conditions
By Elaina Hancock.
As humans continue to send large quantities of carbon into the atmosphere, much of that carbon is absorbed by the ocean, and UConn researchers have found high CO2 concentrations in water can make fish grow smaller.
Researchers Christopher Murray PhD ’19, now at the University of Washington, and UConn Associate Professor of Marine Sciences Hannes Baumann have published their findings in the Public Library of Science (PLoS One).
https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0235817
“The ocean takes up quite a bit of CO2. Estimates are that it takes up about one-third to one-half of all CO2 emissions to date,” says Murray. “It does a fantastic job of buffering the atmosphere but the consequence is ocean acidification.”
Life relies on chemical reactions and even a slight change in pH can impede the normal physiological functions of some marine organisms; therefore, the ocean’s buffering effect may be good for land-dwellers, but not so good for ocean inhabitants.
Baumann explains that in the study of ocean acidification (or OA), researchers have tended to assume fish are too mobile and tolerant of heightened CO2 levels to be adversely impacted.
“Fish are really active, robust animals with fantastic acid/base regulatory capacity,” says Murray. “So when OA was emerging as a major ocean stressor, the assumption was that fish are going to be OK, [since] they are not like bivalves or sea urchins or some of the other animals showing early sensitivities.”
The research needed for drawing such conclusions requires long-term studies that measure potential differences between test conditions. With fish, this is no easy task, says Baumann, largely due to logistical difficulties in rearing fish in laboratory settings.
“For instance, many previous experiments may not have seen the adverse effects on fish growth, because they incidentally have given fish larvae too much food. This is often done to keep these fragile little larvae alive, but the problem is that fish may eat their way out of trouble — they overcompensate – so you come away from your experiment thinking that fish growth is no different under future ocean conditions,” says Baumann.
In other words, if fish are consuming more calories because their bodies are working harder to cope with stressors like high CO2 levels, a large food ration would mask any growth deficits.
Additionally, previous studies that concluded fish are not impacted by high CO2 levels involved long-lived species of commercial interest. Baumann and Murray overcame this hurdle by using a small, shorter-lived fish called the Atlantic silverside so they could study the fish across its life cycle. They conducted several independent experiments over the course of three years. The fish were reared under controlled conditions from the moment the eggs were fertilized until they were about 4 months old to see if there were cumulative effects of living in higher CO2 conditions.
Murray explains, “We tested two CO2 levels, present-day levels and the maximum level of CO2 we would see in the ocean in 300 years under a worst-case emissions scenario. The caveat to that is that silversides spawn and develop as larvae and early juveniles in coastal systems that are prone to biochemical swings in CO2 and therefore the fish are well-adapted to these swings.”
The maximum CO2 level applied in the experiments is one aspect that makes this research novel, says Murray,
“That is another important difference between our study and other studies that focus on long-term effects; almost all studies to date have used a lower CO2 level that corresponds with predictions for the global ocean at the end of this century, while we applied this maximum level. So it is not surprising that other studies that used longer-lived animals during relatively short durations have not really found any effects. We used levels that are relevant for the environment where our experimental species actually occurs.”
Baumann and Murray hypothesized that there would be small, yet cumulative, effects to measure. They also expected fish living in sub-ideal temperatures would experience more stress related to the high CO2 concentrations and that female fish would experience the greatest growth deficits.
The researchers also used the opportunity to study if there were sex-determination impacts on the population in the varying CO2 conditions. Sex-determination in Atlantic silversides depends on temperature, but the influence of seawater pH is unknown. In some freshwater fish, low pH conditions produce more males in the population. However, they did not find any evidence of the high CO2 levels impacting sex differentiation in the population. And the growth males and females appeared to be equally affected by high CO2.
“What we found is a pretty consistent response in that if you rear these fish under ideal conditions and feed them pretty controlled amounts of food, not over-feeding them, high CO2 conditions do reduce their growth in measurable amounts,” says Murray.
They found a growth deficit of between five and ten percent, which Murray says amounts to only a few millimeters overall, but the results are consistent. The fish living at less ideal temperatures and more CO2 experienced greater reductions in growth.
Murray concludes that by addressing potential shortcomings of previous studies, the data are clear: “Previous studies have probably underestimated the effects on fish growth. What our paper is demonstrating is that indeed if you expose these fish to high CO2 for a significant part of their life cycle, there is a measurable reduction in their growth. This is the most important finding of the paper.”
This work was funded by the National Science Foundation grant number OCE #1536165. You can follow the researchers on Twitter @baumannlab1 and @CMurray187.
https://today.uconn.edu/2020/08/uconn-research-carbon-ocean-can-lead-smaller-fish/